Molecular Mechanics Interoperable Components (MMIC)
Molecular Mechanics (MM) is deceptively difficult to execute at any level of experience. Even self-proclaimed experts in MM will say that when they “do MM” or “do Molecular Dynamics (MD),” they are actually spending most of their time on everything except running the simulations and analysis. Most of their time is spent on all of the required secondary skills such as wrapping their heads around the myriad of methods and techniques required to execute MM/MD such as basic simulation settings like temperature control or time propagation; sifting through sampling enhancements to reduce how long simulations take to yield an accurate calculation; slogging through translating ill defined, or incorrectly formatted input files and system preparation to provide the simulation; choosing analysis schemes to process the data yielded by whatever software they are running; or even how to efficiently run code on existing or remote hardware. Each of these choices all preclude any specialty techniques or experiments, and all of these factors complicate even the simplest of MM efforts to the point of needing several PhD’s just in the algorithms to understand what is happening. It may seem that the MM field is unapproachable or at minimum discourages experimentation.
The MM Interoperable Components (MMIC) project at MolSSI seeks to reduce the barrier to entry, allow rapid dissemination of bleeding edge codes, and promote experimentation in the MM and MD spaces. MMIC is an interoperability schema for common workflow components, providing a way to easily share implementations. This schema defines how structured data enters and exits these components through a structured set of I/O pipes. The internals of these components are arbitrary and left up to the design of the end-user themselves. The schema and the components are agnostic to any workflow or even script which is wrapped around them and does not define the context under where or on what hardware it can be used beyond what its I/O defines. Components therefore are valid programmatically from the context of MMIC if they accept the defined inputs and return the defined outputs. The scientific validity of the outputs, however, is left to the individual application of a set of components.
Components are defined at a high scientific level, closer to real world physics and chemistry books, and away from the concepts unique to computer and software definitions of what chemistry, molecular systems, and measurable observables are. This way it is easily understood what each component is aiming to do, rather than obfuscate behind program-specific terminology. In doing so, MMIC’s components also force developers to encapsulate their expertise into the code, making not only the components portable, but able to be experimented against one another. Having expertise encapsulated means non-expert users, those just looking to quickly setup MM experiments, or even those with no computational experience, can design and deploy entire pipelines with minimal effort.
Importantly, this will be software agnostic at a theory level, it is only an API. E.g. The schema specification could be written in Python Abstract Functions, JSON Schema-like, or C++ headers. In practice, these will be codified as an API and implemented in Python. The MMIC project will build reusable components that can be used either standalone or by a number of workflow tools. We are actively avoiding becoming “Yet Another Workflow System” [arXiv 1905.11863]
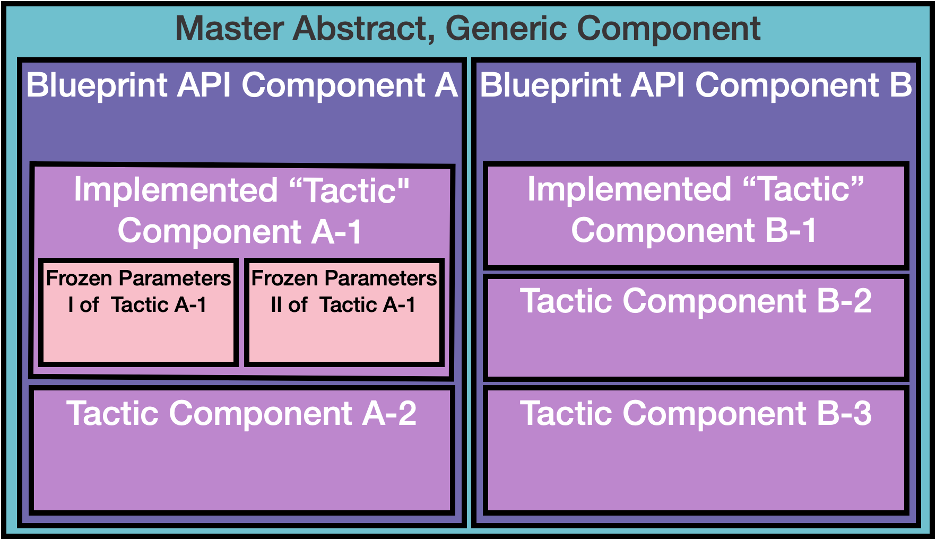
Simply defining this schema is insufficient for adoption, or propagation of these components. MolSSI will facilitate a means for caching and fetching implementations of the components, as submitted by the community. MolSSI is developing practical components in house and deploying them so the communities have something to build on. Component implementation will be hierarchical (see image) such that each implementation comes from a common abstract API level, an example of this shown in the image above. The longer term plan is to cache an index of the implemented components through a tagging system, and store them inside DOI labeled containers on Docker Hub or Singularity Hub and potentially run them through Kubernetes. This will allow for static, self-contained components which can be inserted into any existing pipeline.
You can find more information on MMIC website.
